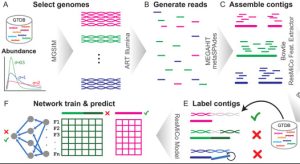
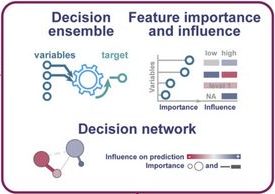
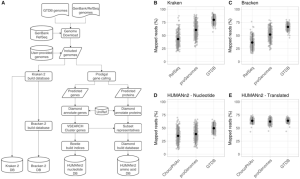
StrainVis: interactive visual strain-level analysis of microbiome data
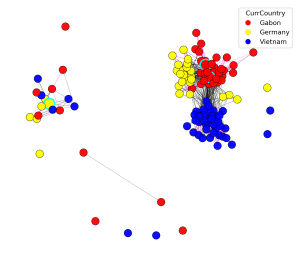
StrainVis is a Python-based web application for visual analyses and interactive exploration of the results obtained by the SynTracker pipeline or by other strain tracking methods, based on ANI (Average Nucleotide Identity). StrainVis accepts either SynTracker’s output file ‘synteny_scores_per_region.csv’ or an ANI file, obtained by another method, containing either one reference genome or multiple reference genomes (usually, one reference genome per species). It presents accordingly analyses for each species separately and for multiple species together. A metadata file (matches…