StrainVis is a Python-based web application for visual analyses and interactive exploration of the results obtained by the SynTracker pipeline or by other strain tracking methods, based on ANI (Average Nucleotide Identity).
StrainVis accepts either SynTracker’s output file ‘synteny_scores_per_region.csv’ or an ANI file, obtained by another method, containing either one reference genome or multiple reference genomes (usually, one reference genome per species). It presents accordingly analyses for each species separately and for multiple species together.
A metadata file (matches to the samples that were previously compared by SynTracker) may also be provided in order to enable a deeper analysis of SynTracker’s results.
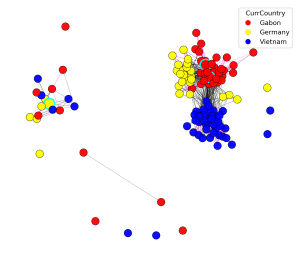
StrainVis allows the user to interactively select the presented plots and to change visual parameters in each plot. It also enables to interactively select the metadata feature by which the samples should be grouped and coloured (for each plot separately).
Each of the presented plots can be downloaded and saved as a high-resolution image, in several common image formats.