We present StrainVis, a new web-based tool designed to simplify and unify strain-level analysis of microbiome data.
Microbial species include multiple strains differing in their genotype, as a result of accumulation of both single nucleotide variants (SNVs) and larger structural genomic changes. While computational tools exist to capture each of these signals, combining them remains technically challenging and typically requires advanced bioinformatic expertise.
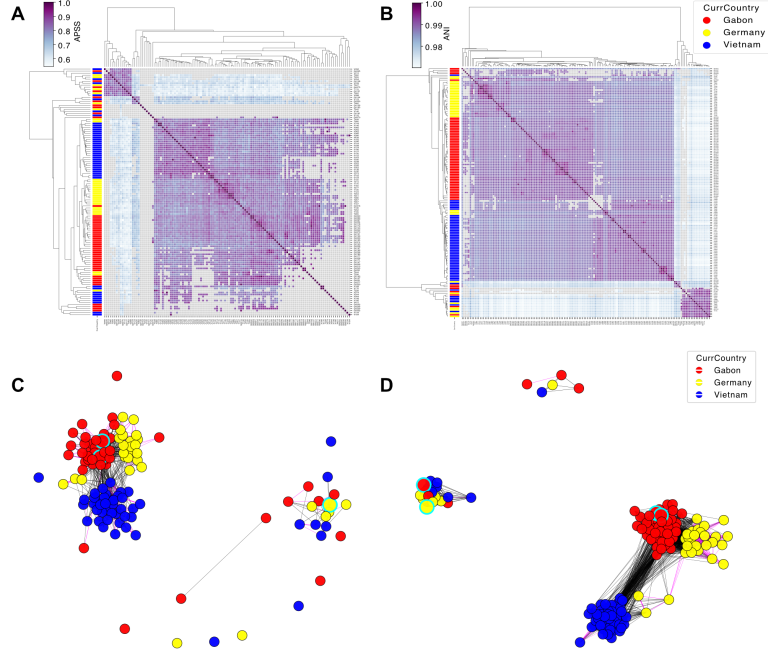
StrainVis bridges this gap by integrating outputs from both structure-based (synteny/APSS, i.e., the output of our tool SynTracker) and SNV-based (ANI) approaches into a single, interactive, web-based interface. The platform enables per-species and multi-species comparisons, incorporation of metadata and gene annotations, and generation of statistical summaries and publication-ready visualizations—all without having to write a single line of code, or have any previous bioinformatic knowledge.
By lowering technical barriers and enabling joint interpretation of complementary signals of genomic variation, StrainVis opens the door for broader adoption of high-resolution, strain-level microbiome analyses.
You can download StrainVis and learn more about the tool at: https://github.com/leylabmpi/StrainVis
Read the full paper: https://www.biorxiv.org/content/10.64898/2026.03.11.711087v1